| Issue |
J. Eur. Opt. Society-Rapid Publ.
Volume 22, Number 1, 2026
|
|
|---|---|---|
| Article Number | 10 | |
| Number of page(s) | 13 | |
| DOI | https://doi.org/10.1051/jeos/2026005 | |
| Published online | 20 February 2026 | |
Review Article
Research on biological detection technology based on fluorescence principles
1
Shandong Laboratory of Advanced Biomaterials and Medical Devices in Weihai, Shandong 264200, PR China
2
Suzhou Institute of Biomedical Engineering and Technology, Chinese Academy of Science, Suzhou 215163, PR China
* Corresponding authors: This email address is being protected from spambots. You need JavaScript enabled to view it.
(Maojie Jiang); This email address is being protected from spambots. You need JavaScript enabled to view it.
(Fuqiang Ma)
Received:
16
December
2025
Accepted:
16
January
2026
Abstract
Biological detection technology plays an important role in the fields of disease diagnosis, environmental monitoring and food safety. Among them, detection technology based on optical principle has the advantages of high sensitivity, fast response and non-destructive, which has led to its rapid development. In this review, we systematically review the principles, progress and applications of fluorescence detection, which achieves ultrasensitive detection and dynamic monitoring of biomolecules through the strategies of fluorescence labeling, fluorescence lifetime imaging, fluorescence polarization immunoassay and fluorescence resonance energy transfer. In the future, the combination of multiple technologies, intelligent data analysis, and portable device development will become important directions for the development of optical detection technology, further promoting its application in precision medicine and environmental monitoring.
Key words: Fluorescence detection / Fluorescent labeling / Fluorescence lifetime imaging microscopy / Fluorescence polarization immunoassay / Fluorescence resonance energy transfer
© The Author(s), published by EDP Sciences, 2026
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
1 Introduction
With the deepening of life science research and the enhancement of health needs, bioassay technology has demonstrated significant application value in key areas such as disease diagnosis, drug discovery and development, environmental monitoring and food safety. Although the capability of biological detection technology has been greatly improved with the development of various technologies, researchers still need to face several key scientific issues and technological challenges: (1) The intrinsic complexity of biological samples (e.g., blood, tissue) introduces significant background noise, non-specific adsorption and matrix effects in samples seriously affect the accuracy and reproducibility of analysis results. Developing efficient matrix management strategies has become the key to improving detection sensitivity [1]; (2) The insufficient ability of multi detection makes it difficult for existing technologies to synchronously achieve multi index analysis, which seriously limits its application in systems biology research [2, 3]; (3) The gap between on-site testing needs and laboratory performance, as well as the lack of a standardization system, constrains the clinical translation and industrial promotion of the technology [4].
With its unique advantages (high sensitivity, fast response, non-destructive and real-time monitoring capability) [5–8], optical detection technology has become a core driving force for the development of bio-detection technology. The development of fluorescent tools, particularly the emergence of various fluorescent sensors, has brought revolutionary breakthroughs in our ability to deeply explore biological dynamic processes at the cellular level. By integrating probes such as small-molecule dyes and fluorescent proteins, these sensors have successfully enabled real-time monitoring and quantitative analysis of cellular characteristics, including proteins, RNA, DNA, small molecules, as well as pH and membrane potential [9]. In the field of medical diagnosis, the breakthrough progress of optical detection technology has provided a revolutionary tool for early screening and accurate diagnosis and treatment of major diseases. Liquid biopsy technology represented by circulating tumor DNA detection, for example, has realized a major leap from theory to clinical application of early cancer diagnosis through ultra-high sensitivity optical detection method [10–12]. In food safety monitoring, optical biosensing technology provides innovative solutions to key quality control issues such as pesticide residues. Rapid testing platforms based on optical principles not only meet the needs of consumer health protection, but also provide technical support for the sustainable development of agriculture and the assessment of compliance with international trade standards [13]. In the field of environmental monitoring, the new optical biosensor has successfully realized high-throughput and real-time monitoring of water pollutants (such as organophosphorus pesticides), and its detection performance is significantly better than the traditional enzyme inhibition method [14, 15].
As a cornerstone method in the field of optical biosensing, fluorescence detection technology occupies an irreplaceable position in life science research and clinical diagnosis due to its excellent sensitivity, excellent molecular specificity and diverse signal transduction mechanisms [16] Through the coordinated development of multi-dimensional detection strategies such as fluorescence labeling technology, fluorescence lifetime imaging (FLIM), fluorescence polarization immunoassay (FPIA) and fluorescence resonance energy transfer (FRET), the technology system has built biological detection, environmental detection and other platforms. Among them, the development of new fluorescent markers such as near-infrared fluorophores and ratiometric probes has significantly improved the selectivity and signal-to-noise ratio of detection [17, 18]; Fluorescence lifetime imaging technology effectively overcomes autofluorescence interference in complex biological samples by resolving nanosecond fluorescence lifetime differences [19, 20]; Fluorescence polarization technology uses the dynamic changes of molecular rotational relaxation time to provide a unique analytical perspective for the study of molecular interactions [21]; Owing to its exceptional sensitivity to nanoscale proximity (1–10 nm), fluorescence resonance energy transfer serves as a powerful “molecular ruler”. This capability makes it an indispensable technique for revealing protein-protein interactions and monitoring nucleic acid conformational changes in real time [22], and is in great demand in emerging fields such as genomics and proteomics [23]. These technical advantages enable fluorescence detection methods to achieve rapid, high-throughput, ultra-high sensitivity and specific detection, which perfectly fits the urgent needs of modern genomics and proteomics research for precise analytical tools [24]. Multi-color fluorescence detection enables the simultaneous detection of multiple biomarkers for a more comprehensive understanding of the condition, leading to more accurate medical diagnosis [25]. This multiple detection ability provides more comprehensive biological information for disease diagnosis. For example, through the simultaneous detection of multiple tumor markers, the accuracy and reliability of cancer diagnosis can be significantly improved [26]. However, fluorescence technology is still limited by problems such as photobleaching and autofluorescence interference, driving the exploration of novel probes and detection modes.
2 Fluorescence detection
Fluorophores are a class of chromophores that can specifically respond to photoexcitation and emit fluorescence, and their luminescence mechanism is shown in Figure 1 [27], where electrons jump to the singlet excited state (S1 or higher energy level) when the ground state molecule (singlet state S0) absorbs photons of a specific wavelength. Due to the instability of the excited state, the molecule rapidly relaxes to the lowest vibrational energy level of S1 through nonradiative relaxation (such as internal transitions), and then returns to the ground state through radiative leaps and releases photons with energies lower than that of the excitation light, resulting in a redshift of the emission wavelength, a phenomenon known as the Stokes shift. As photosensitive compounds, fluorescent dyes have characteristic absorption (excitation spectrum) and emission spectra determined by their electronic structure. Due to their ability to efficiently convert short-wavelength light energy into long-wavelength fluorescence, fluorescent dyes have been widely used as highly sensitive optical probes in the fields of biomarking, cellular imaging and tissue detection. Building on this foundation, this technology enables the quantitative, real-time observation of biomolecules with exceptional sensitivity, utilizing a suite of techniques such as fluorescent labeling, FLIM, FPIA, and FRET.
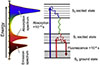 |
Fig. 1 Energy levels and radiative transition from the excited state (S1/S2) to the ground state (S0). |
2.1 Fluorescent labeling technique
Fluorescent labeling technique is a method for qualitative and quantitative analysis of the measured components through physical adsorption or chemical coupling of fluorescent substances with the target molecules specifically combined to form a fluorescent labeling complex. The core component of biofluorescent probes is a class of compounds that can absorb light at specific wavelengths and emit fluorescence at longer wavelengths. This technique can be traced back to the antibody fluorescence labeling studies in the 1940s [28, 29], and has now developed into an important analytical tool with significant advantages such as fast detection speed [30], high sensitivity [31], low sample consumption and non-radioactivity [32], and has been widely used in the fields of protein localization and activity detection, biomolecular interaction studies, and in vivo dynamic process monitoring. To address the poor selectivity of natural fluorescent dyes, researchers have significantly improved the selective recognition ability of probes by coupling fluorophores with biological targeting molecules such as antibodies [33], nucleic acid aptamers [34], and developing fluorescein derivatives [35, 36], which not only provides a powerful tool for studying biomolecular dynamics and photophysical properties, but also shows important application value in biomedical fields (such as anticancer drug research) [37].
A long-standing challenge in the O-GlcNAc field has been the lack of tools for specific, non-antibody-based detection. Traditional immunoassays, while widely used, often suffer from cross-reactivity and limited ability to distinguish O-GlcNAcylation from other post-translational modifications in complex samples. To address this specificity gap, the breakthrough work by Wu et al. [38] introduced a tandem glycan labeling strategy. This method converts the O-GlcNAc moiety into a fluorescently tagged epitope through sequential enzymatic steps, thereby enabling highly specific and sensitive detection independent of conventional antibodies, as shown in Figure 2a. This method achieves two-step labeling by galactosylating O-GlcNAc by B4GalT1 and then connecting the fluorescent moiety by ST6Gal1-mediated sialylation. The experimental validation showed that the method could efficiently detect O-GlcNAc modification in HEK293 nuclear extract and recombinant proteins (such as CK2, HIF1 α), and the signal was specific after pre-treatment with OGA/OGT. Compared with the traditional antibody method, this method avoids the side reactions of click chemistry, has a sensitivity of femtomolar level, and supports direct imaging by SDS-PAGE. This technology provides a new tool for the visualization and functional study of low abundance O-GlcNAc protein, reveals the potential O-GlcNAc regulatory mechanism of HIF1 α and other targets, and opens up a new way for glycobiology research.
 |
Fig. 2 (a) Fluorescent detection of O‑GlcNAc via Tandem Glycan Labeling. (b) Fluorescent labeling technique for lipid and surfactant preformed vesicles. (c) Efficient labeling of vesicles with lipophilic fluorescent dyes via the Salt-Change method. |
In the field of extracellular vesicles (EVs), fluorescent labeling technology also plays an important role. Mousseau et al. [39] developed a universal fluorescent labeling technology based on the fat soluble carbocyanine dye PKH67 for the tracking study of prefabricated liposomes and surfactant vesicles, as shown in Figure 2b. This method avoids fluorescence quenching caused by dye aggregation by optimizing the molar ratio of dye to lipid, and does not require complex purification steps. The experimental verification showed that this technology could efficiently label neutral phospholipids (DPPC), negatively charged pulmonary surfactant (Curosurf) and positively charged double chain surfactant (TEQ) vesicles, and the size and surface potential of the vesicles remained stable after labeling. Through fluorescence microscopy and confocal imaging, the research team successfully tracked the interaction between nanoparticles and vesicles and their separation process in cells, visualized the adsorption behavior of cationic surfactants on cotton fibers, and revealed the characteristics of their deposition in the form of bilayers or vesicles. This technology provides a simple and efficient solution for vesicle tracking in biomedical and industrial applications.
However, conventional methods are often inefficient during labeling due to the low water solubility of the dyes, and the use of high concentrations of dyes is prone to problems such as nonspecific labeling and changes in vesicle size. To address these challenges, Cha et al. [40] proposed a vesicle fluorescence labeling technique based on salt concentration modulation, the Salt-Change Method, as shown in Figure 2c. The labeling efficiency of the lipophilic dye DiI was significantly enhanced by dynamically adjusting the NaCl concentration (low-salt condition promotes dye dispersion, and high-salt condition induces free dye aggregation) combined with filter membrane purification. Experimental validation showed that the method is applicable to cell-derived vesicles (CDVs), natural EVs, and synthetic liposomes, and that the size distribution and membrane protein function (such as antibody binding to CD63/PD-1) of the labeled vesicles remain intact. In addition, the method is compatible with a wide range of dyes (DiD, PKH67), is easy to perform, and reduces nonspecific labeling by lowering the dye concentration. This technique provides a highly sensitive and specific solution for the study of EVs at the single-particle level and has significant potential for application in liquid biopsy and nanomedicine.
Fluorescent labeling technology has become an important tool in biomedical research. From the specific detection of glycosylation modification to the optimization of vesicle tracing technology, these innovative achievements not only expand the application scope of fluorescent labeling technology, but also provide a powerful tool for in-depth research in related fields. With the emergence of new fluorescent probes and labeling strategies, fluorescent labeling technology will continue to provide more accurate and efficient solutions for life science research and promote the in-depth development of related fields.
2.2 Fluorescence lifetime imaging microscopy
With the aid of appropriate biomarkers and transfection strategies, fluorescence imaging technology provides access to multidimensional information – such as three-dimensional spatial localization, spectral signatures, and polarization states. However, traditional fluorescence intensity imaging is susceptible to interference from many factors, including fluctuation of excitation light source, probe concentration difference, photobleaching effect, and changes in instrument parameters (such as focal length and detector gain) [41, 42]. Fluorescence lifetime imaging microscope (FLIM) significantly improves the accuracy and information dimension of biological sample analysis by detecting the spatio-temporal characteristics of the decay of excited states of fluorophores. It is important to note that while the fluorescence lifetime in FLIM is not absolutely invariant, its dependence on factors like probe concentration and excitation light intensity is orders of magnitude weaker than that of fluorescence intensity. Furthermore, it exhibits superior stability against photobleaching. This robustness establishes lifetime as a highly reliable parameter for quantitatively sensing changes in the local microenvironment [43]. In living cell research, FLIM can quantitatively resolve pH gradient (range 6.0–8.0), calcium ion concentration (detection limit up to nm level) and intermolecular distance (calculated by Förster resonance energy transfer efficiency), and its time resolution can reach picosecond level [44]. This technology shows unique value in dynamic monitoring of protein interactions, metabolic state transition and other key physiological processes [45]. A prime application of FLIM is the quantification of metabolic heterogeneity in cell populations. This is achieved by monitoring the fluorescence lifetime decay of endogenous metabolic co-factors, such as NADH and FAD. Unlike intensity-based measurements, the lifetime signal is independent of fluorophore concentration, allowing FLIM to eliminate concentration-dependent bias and provide a more accurate readout of cellular metabolic states [46, 47]. This non-invasive single-cell metabolic analysis capability provides a new technological paradigm for precision medicine and drug screening. At present, FLIM technology has been combined with multiphoton microscopy and super-resolution imaging, further expanding its application boundaries in the dynamic monitoring of subcellular organelles and the study of nanoscale molecular interactions [48].
FLIM technology provides a breakthrough tool for the study of the dynamics of G-quadruplex DNA (G4) in living cells by detecting the spatial and temporal distribution of fluorophore excited state lifetimes. Summers et al. [49] developed a FLIM-based dynamic detection of G4 in living cells, as shown in Figure 3a, using a DAOTA-M2 probe to achieve specific recognition through fluorescence lifetime differences (9–12 ns in the G4-bound state vs. 5–7 ns in the double-stranded DNA-bound state). This technique breaks through the limitations of traditional antibody labelling and quantitatively assesses the interaction of G4 with small molecules and metal-salphen complexes (Ni > VO > Cu) using fluorescence lifetime displacement analysis (FLIDA), with a high degree of intracellular core-binding potency in line with in vitro data. It was confirmed that reduced expression of the deconjugating enzyme FancJ/RTEL1 prolongs DAOTA-M2 lifespan, suggesting increased G4 stability. This technique provides a new approach for the study of nucleic acid structure in living cells and G4-targeted drug development.
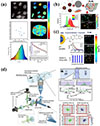 |
Fig. 3 (a) Visualising G-quadruplex DNA dynamics in live cells by fluorescence lifetime imaging microscopy. (b) Functionalized CdSe/ZnS quantum dots for intracellular pH measurements by fluorescence lifetime imaging microscopy. (c) A novel quantum dot-based pH probe for long-term fluorescence lifetime imaging microscopy experiments in living cells. (d) High-throughput fluorescence lifetime imaging flow cytometry. |
The quantitative analysis of intracellular pH benefits uniquely from FLIM. Because the fluorescence lifetime of specific probes is directly modulated by proton concentration, and crucially, is independent of probe concentration, FLIM provides a robust and artifact-free method for mapping pH with high precision across cells and organelles. Pacheco-Liñán et al. [50] developed three functionalized CdSe/ZnS quantum dot (QD) probes (CdSe/ZnS-p, CdSe/ZnS-pE, and CdSe/ZnS-pK), and it was found that the fluorescence lifetimes of these nanoprobes exhibited a significant response to pH variations by steady-state and time-resolved fluorescence spectroscopy, as shown in Figure 3b. Theoretical calculations indicate that the intramolecular interaction of the protonated amine group with the sulfur atoms of the ZnS shell layer is the key mechanism for pH sensitivity: this interaction leads to fluorescence bursting under acidic conditions, whereas the ligand deprotonation under alkaline conditions leads to fluorescence enhancement. Cellular experiments confirm that these QD probes are efficiently internalised and distributed in the cytoplasm with low cytotoxicity. Using FLIM technology, researchers have achieved for the first time QD-based absolute measurement of pH in living cells. These probes are able to distinguish different cell types by fluorescence lifetime differences in co-culture experiments, demonstrating their unique advantages in cellular microenvironmental studies. This study provides an important reference for the development of novel FLIM biosensors and promotes the application of fluorescence lifetime imaging technology in the biomedical field.
Based on the previous development of D-penicillamine-functionalized CdSe/ZnS QD probe [50], Pacheco-Liñán et al. further optimized and designed a novel D-penicillamine-histidine peptide-functionalized QD (CdSe/ZnS-pH) probe [51], as shown in Figure 3c. Compared with its predecessor, the new probe demonstrated significant advantages: (1) extended fluorescence lifetime and improved signal-to-noise ratio; (2) extended pH response range to 3–7 and improved sensitivity to 2.42 ns/pH; (3) maintained linear response in a simulated intracellular environment, which overcame signal fluctuations of its predecessor in complex environments; and (4) cellular experiments confirmed its excellent 24-hour stability, whereas the previous generation probe suffered from signal attenuation in long-term experiments. Mechanistic analyses showed that histidine residues facilitated the escape of the probe into the cytoplasm to avoid lysosomal degradation through enhanced pH sensitivity and the “proton sponge hypothesis”. This improvement makes the CdSe/ZnS-pH probe a more reliable tool for long-term pH monitoring in living cells, and provides a new method for cell metabolism research and drug screening.
FLIM Flow Cytometry enables high-throughput, high-resolution analysis of single cells by combining FLIM with flow cytometry. While conventional flow cytometry relies on fluorescence intensity, which is susceptible to interference from concentration, scattering and other factors, FLIM technology takes advantage of the stability of fluorescence lifetimes to dramatically improve detection accuracy. The high-throughput FLIM system developed by Kanno et al. [52] employs a dual-intensity-modulated, continuous-wave beam array to acquire fluorescence lifetimes images at a rate of more than 10,000 cells/second, as shown in Figure 3d [53]. Crucially, this system maintains cell-level resolution, associating each lifetime measurement with an individual cellular event, unlike bulk measurements that average signals across populations. The system successfully distinguished subpopulations of rat glioma cells and captured anticancer drug-induced changes in cell nuclear dynamics, revealing heterogeneous features that are difficult to observe by conventional methods. In addition, the sensitivity of FLIM technology to environmental factors (such as pH, ion concentration) provides a new tool for drug screening and tumour microenvironment studies. The breakthrough of this technology opens up new avenues for cell function analysis and clinical diagnosis.
These research fully demonstrate the broad application prospect of FLIM technology in many biomedical research fields. With probe optimization and system integration (such as quantum dot probes, multimodal imaging, etc.), FLIM has been widely used in live cell dynamic observation, drug screening and clinical diagnosis. In the future, FLIM will play an increasingly important role in precision medicine, drug development and other fields.
2.3 Fluorescence polarization immunoassay
Fluorescence intensity is susceptible to external noise, affecting the sensitivity and reliability of the analysis, whereas rapid and specific diagnosis is essential for the treatment and prevention of infectious diseases [54, 55]. Compared with traditional fluorescence detection, Fluorescence Polarization Immunoassay (FPIA) has significant anti-interference advantages and is a highly sensitive analytical technique based on the rotational kinetic properties of fluorescently labelled molecules. Figure 4 shows a schematic diagram of the principle of Fluorescence Polarization [56]. The core principle of this technology lies in the fact that when a fluorescently labelled molecule is excited by linearly polarised light, the Polarization state of the emitted light is closely related to the rotational state of the molecule, and since the rate of rotational diffusion of a molecule is inversely proportional to its molecular weight, the change of the fluorescence Polarization signal directly reflects the change in molecular weight of the molecule in the process of binding or dissociation [57, 58]. Specifically, when a small molecule fluorescent tracer binds to a large molecule (such as antibody), its rotational speed decreases, leading to an increase in the Polarization of the emitted light; conversely, a decrease in molecular weight during processes such as enzymatic cleavage will manifest itself as a decrease in Polarization [59]. The system employs a ratiometric measurement mode to eliminate common-mode noise such as light source fluctuations by detecting the ratio of fluorescence intensity in parallel and perpendicular directions (I∥/I⊥). Especially when coupled with quench-free reporter probes, it not only reduces the cost of detection, but also effectively suppresses the interference of ambient light and electronic noise, and significantly improves the stability and reliability of the detection [60]. This quantitative relationship enables fluorescence Polarization techniques to monitor the dynamics of biomolecular interactions in real time, and the binding ratios can be extrapolated from the changes in the Polarization values. Based on the above advantages, fluorescence Polarization immunoassay has been widely used in various fields: in basic research for protein-ligand interaction analysis and enzyme kinetic studies [61–63]; in drug discovery and development for the primary screening stage of high-throughput screening [64]; and in clinical diagnostics for rapid detection of infectious disease markers [65]. In addition, the method has shown unique value in biosafety monitoring (such as toxin detection) [66], as it is particularly suited for rapid detection in the field due to the absence of a separation step. With advances in polarised optical components and fluorescent probe technology, the range of applications for this technique continues to expand.
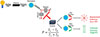 |
Fig. 4 Schematic diagram of fluorescence polarization principle. |
In the field of environmental pollutant detection, FPIA has become an ideal choice for rapid screening because of its simple operation and no complex pre-treatment. Raysyan et al. [67] successfully optimized the FPIA method and detected diclofenac (DCF) in wastewater by designing novel fluorescent markers (including homologous and heterologous structures), as shown in Figure 5a. Research has shown that the FPIA system using DCF-Ahx-AMF markers exhibits excellent selectivity and sensitivity, with detection performance highly consistent with LC-MS/MS methods, and analysis time significantly shorter than traditional ELISA. This work further confirms the low cross reactivity of FPIA to DCF metabolites, highlighting its reliability in complex matrices such as wastewater, providing an important technological paradigm for efficient monitoring of small molecule pollutants in the environment, and promoting the application and development of FPIA in the field of environmental analysis.
 |
Fig. 5 (a) Schematic diagram of fluorescence polarization immunoassay for determining diclofenac in wastewater. (b) Schematic diagram of homogeneous fluorescence polarization immunoassay for rapid determination of erythromycin in milk. (c) Schematic diagram of one-step FPIA detection of maduramycin and salinomycin based on bispecific monoclonal antibodies. |
FPIA also has significant advantages in the field of food safety detection. A FPIA-based homogeneous assay was developed by Duan et al. [68] for the rapid determination of erythromycin (ERY) residues in milk, as shown in Figure 5b. The method was developed by synthesizing five fluorescently labelled tracers (ERY-CMO-4′-AMF, ERM-FITC, ERM-DTAF, ERM-SRSC, ERM-AF647) and pairing them with three monoclonal antibodies (5B2, 6C2, and 6D9), and the optimal combination of monoclonal antibodies, 5B2/ERM-FITC, was compared and screened. By optimizing the reaction conditions (including tracer fluorescence intensity, pH value and antibody concentration, etc.), the method can complete the detection within 5 min, significantly improving the analysis efficiency. The experimental results show that the method has high sensitivity and accuracy for the detection of ERY in milk, and its recovery and precision meet international standards, which can meet the requirements of China and the European Union for the detection of ERY residues. This study provides a fast and reliable solution for food safety monitoring.
In addition, FPIA has also made important progress in multi-residue detection. Huang et al. [69] developed a bispecific monoclonal antibody (BsMAb)-based FPIA method for the simultaneous detection of maduramicin (MAD) and salinomycin (SAL) in chicken meat, as shown in Figure 5c. In this study, BsMAb was obtained by hybridoma technique and combined with specific fluorescent probe (MAD-FITC/SAL-EDF) to establish an efficient detection system, which could be completed in only 10 min with high sensitivity and no cross-reaction. Molecular docking revealed the specific binding mechanism of BsMAb, and the recoveries of the actual samples were 88.7–113.7%, which were in high agreement with the LC-MS/MS results. This study provides an efficient solution for the detection of antibiotic multiresidues in food and fully demonstrates the potential of FPIA technology.
FPIA has shown important application value in many fields with its advantages of high sensitivity, strong anti-interference ability and simple operation. In environmental monitoring, FPIA can efficiently detect pollutants in wastewater. In the field of food safety [70], this method can quickly screen erythromycin in milk and a variety of antibiotic residues in chicken. In addition, FPIA does not need complex separation steps, and is suitable for on-site rapid detection, providing a reliable tool for infectious disease diagnosis, drug screening and other research. With the progress of polarization optical elements and fluorescent probe technology, the application scope of this technology is still expanding.
2.4 Fluorescence resonance energy transfer
In 1948, Theodor Förster first proposed [71] the theory of fluorescence resonance energy transfer (FRET), which describes a non-radiative energy transfer phenomenon: when the distance between the excited state molecule (donor) and the ground state molecule (acceptor) is less than 10 nm, the donor can transfer energy to the acceptor through near-field coupling, which is often referred to as “fluorescence resonance energy transfer” when both the donor and the acceptor are fluorophores. In the process, the donor molecule can return to the ground state through two pathways: emitting fluorescent photons or transferring the energy through FRET [72], as shown in Figure 6 [73]. The existence of FRET adds new pathways for energy dissipation in the donor, leading to a shortening of its excited state lifetime, a property that plays a key role in unravelling the mechanism of interactions between biomolecules. In 1967, the FRET theory was experimentally validated for the first time. Its unique distance dependence (1–10 nm) makes it an ideal ‘optical ruler’ for molecular scale interactions and provides a breakthrough method for quantitative study of biomolecular interactions [74]. In the 1980s, FRET technology was successfully applied to protein conformation studies, and through the combination with other analytical techniques, it has driven the innovation of molecular detection methods [75, 76]. With the development of technology, FRET-based biosensors have gradually become a research hotspot. Their design centers on coupling the donor and acceptor fluorophores within a single, responsive molecular scaffold. This is typically achieved by genetically fusing fluorescent protein pairs (e.g., CFP/YFP) to a sensing domain that undergoes a target-induced conformational change, or by conjugating organic dyes to complementary biomolecular recognition elements (e.g., antibodies, oligonucleotides). When the target analyte binds or the enzymatic reaction occurs, it alters the distance and/or orientation between the donor and acceptor. This change modulates the FRET efficiency, which is directly read out as a quantifiable shift in the fluorescence emission ratio of the acceptor to the donor, enabling sensitive and spatiotemporally resolved detection. This design not only achieves specific detection of molecular interactions, but also dynamically reflects changes in the microenvironment, providing an important tool for molecular mechanism research and biomedical applications.
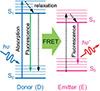 |
Fig. 6 Schematic diagram of fluorescence resonance energy transfer. |
FRET technology has made significant progress in the application of in vivo imaging, especially in tumor metabolism and drug delivery monitoring. Rudkouskaya et al. [77] developed a FRET receptor technology based on the dark bursting agent QC-1, combined with near-infrared Macroscopic Fluorescence Lifetime Förster Resonance Energy Transfer (MFLI-FRET), enabling multiple non-invasive imaging for simultaneous monitoring of tumour glucose metabolism and receptor-ligand binding in living mice, as shown in Figure 7a. This technique enables the simultaneous quantification of two independent biological processes in the same tumor: 1) intracellular drug delivery efficiency, measured by the FRET signal generated when transferrin (Tf) donors and dark-quencher acceptors (e.g., Tf-QC-1) bind in proximity to transferrin receptors (TfR) and undergo internalization; and 2) tumor glycolytic activity, assessed by the fluorescence intensity of a 2-deoxyglucose (2-DG) probe that accumulates upon cellular uptake via glucose transporters (GLUT1). It was found that drug delivery efficiency was lower in highly metabolically active tumour regions, revealing a potential correlation between metabolic status and the efficacy of targeted therapies. Multiscale validation (in vitro microscopy and in vivo wide-field imaging) confirmed the stability of QC-1 as a FRET acceptor. The study realized the synchronous monitoring of drug targeted delivery and metabolic response in living animals for the first time, providing an important tool for optimizing preclinical drug research and development, and also opening up a new way to understand the relationship between tumor microenvironment and therapeutic response.
 |
Fig. 7 (a) Schematic illustration of non-invasive tumor imaging based on multi parameter glucose metabolism and receptor-ligand engagement using dark quenched FRET acceptor. (b) Schematic illustration of Fluonanobody-based nanosensor via fluorescence resonance energy transfer for ultrasensitive detection of ochratoxin A. (c) Schematic diagram of high-performance fluorescent microspheres based on fluorescence resonance energy transfer mode for lateral flow immunoassay. (d) Schematic illustration of the Principle of the Ratiometric Fluorescence Aptasensor Based on Multivalent Aptamer Encoded DNA Flowers. |
In the field of food safety, FRET technology provides an efficient solution for rapid detection. Su et al. [78] developed a Fluonanobody-based nanosensor (FN-Nanosens) for the ultrasensitive detection of ochratoxin A (OTA) based on the FRET technique, as shown in Figure 7b. The sensor uses a fusion protein (SGFP-Nb) of a superfolded green fluorescent protein (SGFP) and a nanobody (Nb) as an energy donor and OTA-labelled quantum dots (OTA-RQDs) as an energy acceptor. The efficient FRET effect was achieved by optimising the reaction conditions (e such as donor-acceptor molar ratio, pH, etc.). Molecular docking analysis showed that the binding of SGFP-Nb to OTA was dependent on hydrogen bonding, hydrophobic interaction, etc., ensuring high selectivity. Compared with the chromatographic method, this method is easy to operate, low cost and suitable for the rapid detection of OTA in food.
The combination of FRET technology and new fluorescent materials further improves the detection performance. Wang et al. [79] successfully constructed FRET modes by encapsulating two aggregation induced emission (AIE) fluorescent groups (TCBPEME and BAPF) with spectral overlap in polymer microspheres, as shown in Figure 7c. In this study, AIE doped polymer microspheres (DAIEPMs) were synthesized by microemulsion method. DAIEPMs exhibit significant large Stokes shift and fluorescence enhancement effects, while possessing high quantum yield and excellent energy transfer efficiency. At the application level, DAIEPMs have been successfully used as fluorescent markers in lateral flow immunoassay (LFIA), achieving ultra-high sensitivity detection of organochlorine pesticide chlorothalonil (CTN), with a detection limit significantly better than traditional quantum dot microsphere and gold nanoparticle labeling methods. In addition, DAIEPMs exhibit excellent anti-interference performance and thermal stability in complex matrices, providing new ideas for optimizing the performance of fluorescent materials. This study not only highlights the potential of FRET technology in improving detection sensitivity, but also lays an important foundation for developing high-performance optical detection platforms.
FRET technology also shows unique advantages in allergen detection. Qi et al. [80] developed a ratiometric fluorescent sensor based on multivalent aptamer-encoded DNA flowers (Mul-DNFs) for the detection of the peanut allergen Ara h1, as shown in Figure 7d. A large number of aptamers were modified on the surface of the self-assembled Mul-DNFs by rolling circle amplification (RCA), which dramatically enhanced the binding ability. The sensor utilises Cy3/Cy5-labelled complementary DNA to construct a FRET platform, and the dissociation of Cy5-cDNA upon Ara h1 binding induces a change in FRET signal for quantitative detection. The recoveries of the actual sample assay reached 95.7-106.3%, providing a new strategy for allergen analysis in complex matrices.
FRET technology has become an important tool for studying molecular interactions due to its nanoscale distance sensitivity and efficient energy transfer characteristics. From in vivo tumor metabolism monitoring, food safety detection, and allergen analysis, FRET technology continues to expand its application boundaries through innovative donor receptor design and multidisciplinary integration. In recent years, with the development of new fluorescent materials (such as AIE polymer microspheres) and nanosensors, the detection sensitivity, anti-interference ability and dynamic monitoring performance of FRET have been significantly improved. In the future, further optimizing the specificity and stability of FRET probe and deepening its application in multimodal imaging and clinical diagnosis will open up broader prospects for life science and medical research.
3 Conclusions and outlook
Although the above technologies have made significant progress, it is often difficult for a single technology to meet the demand for high-throughput and high-precision detection of complex samples. The future development of optical detection technology will show a trend of multi-dimensional innovation, mainly in the following aspects: first, in terms of detection sensitivity, the application of new nanomaterials (such as two-dimensional materials, metal-organic frameworks) and quantum dots will break through the existing detection limits. Secondly, the synergistic integration of microfluidic chip and optical detection will significantly enhance the system response speed and achieve dynamic monitoring with millisecond time resolution. In terms of specificity enhancement, intelligent analysis strategies based on machine learning algorithms, such as deep neural network-assisted Raman spectral resolution, will dramatically improve the recognition accuracy of complex samples. In the future, cross technology integration will become the key direction to improve the detection performance. Fluorescence plasmon combination, such as metal enhanced fluorescence technology, can significantly improve the fluorescence signal intensity and stability by combining the field enhancement effect of LSPR; Chemiluminescence – nanomaterial synergy and nanoenzyme catalyzed chemiluminescence reaction can further reduce the detection limit and expand the ability of multiple detection; AI assisted analysis and machine learning algorithm can be used for rapid analysis of multimodal data and improve the identification accuracy of complex samples. With the cross innovation of technology, the development of detection technology in the future will tend to be “highly sensitive, multimodal, intelligent and portable”. Through multi technology collaboration and engineering design, the existing bottlenecks will be gradually solved, providing stronger tools for life science, clinical diagnosis, public safety and other fields.
Funding
This work was supported by Shandong Provincial Laboratory Project [SYS202209].
Conflicts of interest
The authors have nothing to disclose.
Data availability statement
This article did not generate any new datasets.
Author contribution statement
Conceptualization, Maojie Jiang and Fuqiang Ma; Data curation, Yanna Lin, Peng Yin, Xiangyu Jiang; Formal analysis, Mengjie Huang and Baihui Zhang; Investigation, Maojie Jiang, Yanna Lin and Xuan Fang; Writing – Original Draft Preparatio, Maojie Jiang; Writing – Review and Editing, Maojie Jiang and Fuqiang Ma; Visualization, Maojie Jiang and Peng Yin; Funding acquisition, Fuqiang Ma. All authors have read and agreed to the published version of the manuscript.
Acknowledgments
Fuqiang Ma was supported by Shandong Provincial Laboratory Project [SYS202209].
References
- Chiu ML, Lawi W, Snyder ST, Wong PK, Liao JC, Gau V, Matrix effects – a challenge toward automation of molecular analysis, JALA-J. Lab Autom. 15(3), 233–242 (2010). https://doi.org/10.1016/j.jala.2010.02.001. [Google Scholar]
- Sapsford KE, Bradburne C, Delehanty JB, Medintz IL, Sensors for detecting biological agents, Mater. Today 11(3), 38–49 (2008). https://doi.org/10.1016/S1369-7021(08)70018-X. [CrossRef] [Google Scholar]
- Papakostas GI, Shelton RC, Kinrys G, Henry ME, Bakow BR, Lipkin SH, et al., Assessment of a multi-assay, serum-based biological diagnostic test for major depressive disorder: A pilot and replication study, Mol. Psychiatr. 18(3), 332–339 (2013). https://doi.org/10.1038/mp.2011.166. [Google Scholar]
- Yang SM, Lv S, Zhang W, Cui Y, Microfluidic point-of-care (POC) devices in early diagnosis: A review of opportunities and challenges, Sensors 22(4), 1620 (2022). https://doi.org/10.3390/s22041620. [Google Scholar]
- Noor ASM, Talah A, Rosli MAA, Thirunavakkarasu P, Tamchek N, Increased sensitivity of Au-Pd nanolayer on tapered optical fiber sensor for detecting aqueous ethanol, J. Eur. Opt. Soc.-Rapid Publ. 13(1), 28 (2017). https://doi.org/10.1186/s41476-017-0056-6. [Google Scholar]
- Minkovich VP, Sotsky AB, Tapered photonic crystal fibers coated with ultra-thin films for highly sensitive bio-chemical sensing, J. Eur. Opt. Soc.-Rapid Publ. 15, 7 (2019). https://doi.org/10.1186/s41476-019-0103-6. [Google Scholar]
- Fischer F, Frenner K, Granai M, Fend F, Herkommer A, Data-driven development of sparse multi-spectral sensors for urological tissue differentiation, J. Eur. Opt. Soc. 19(1), 8 (2023). https://doi.org/10.1051/jeos/2023030. [Google Scholar]
- Bouquet G, Kaspersen K, Haugholt KH. Optical measurement instrument for detection of powdery mildew and grey mould in protected crops, J. Eur. Opt. Soc. 20, 1 (2024). https://doi.org/10.1051/jeos/2024024. [Google Scholar]
- Specht EA, Braselmann E, Palmer AE, A Critical and comparative review of fluorescent tools for live-cell imaging, Annu. Rev. Physiol. 79(1), 93–117 (2017). https://doi.org/10.1146/annurev-physiol-022516-034055. [Google Scholar]
- Crowley E, DiNicolantonio F, Loupakis F, Bardelli A, Liquid biopsy: monitoring cancer-genetics in the blood, Nat. Rev. Clin. Oncol. 10(8), 472–484 (2013). https://doi.org/10.1038/nrclinonc.2013.110. [Google Scholar]
- Stoetzer OJ, Fersching DM, Salat C, Steinkohl O, Gabka CJ, Hamann U, et al., Prediction of response to neoadjuvant chemotherapy in breast cancer patients by circulating apoptotic biomarkers nucleosomes, DNAse, cytokeratin-18 fragments and surviving, Cancer Lett. 336(1), 140–148 (2013). https://doi.org/10.1016/j.canlet.2013.04.013. [Google Scholar]
- Kitz J, Lowes LE, Goodale D, Allan AL, Circulating tumor cell analysis in preclinical mouse models of metastasis, Diagnostics 8(2), 30 (2018). https://doi.org/10.3390/diagnostics8020030. [Google Scholar]
- Omarov R, Agarkov A, Rastovarov E, Shlykov S, Modern methods for food safety, Eng. Rural Dev. 24(26.05), 960–963 (2017). https://doi.org/10.22616/ERDev2017.16.N195. [Google Scholar]
- Long F, Zhu A, Shi H, Recent advances in optical biosensors for environmental monitoring and early warning, Sensors 13(10), 13928–13948 (2013). https://doi.org/10.3390/s131013928. [Google Scholar]
- Herrera-Domínguez M, Morales-Luna G, Mahlknecht J, Cheng Q, Aguilar-Hernández I, Ornelas-Soto N, Optical biosensors and their applications for the detection of water pollutants, Biosensors 13(3), 370 (2023). https://doi.org/10.3390/bios13030370. [Google Scholar]
- Mathies RA, Peck K, Stryer L, Optimization of high-sensitivity fluorescence detection, Anal. Chem. 62(17), 1786–1791 (1990). https://doi.org/10.1021/ac00216a012. [Google Scholar]
- Hao Y, Nguyen KH, Zhang Y, Zhang G, Fan S, Li F, et al., A highly selective and ratiometric fluorescent probe for cyanide by rationally altering the susceptible H-atom, Talanta 176, 234–241 (2018). https://doi.org/10.1016/j.talanta.2017.08.032. [Google Scholar]
- Tian X, Murfin LC, Wu L, Lewis SE, James TD, Fluorescent small organic probes for biosensing, Chem. Sci. 12(10), 3406–3426 (2021). https://doi.org/10.1039/D0SC06928K. [Google Scholar]
- Suhling K, Hirvonen LM, Levitt JA, Chung PH, Tregido C, Marois AL, et al., Fluorescence lifetime imaging (FLIM): Basic concepts and recent applications, Springer Ser. Chem. Phys. 111, 119–188 (2015). https://doi.org/10.1007/978-3-319-14929-5_3. [Google Scholar]
- Malý P, Strachotová D, Holoubek A, Heřman P, Interferometric excitation fluorescence lifetime imaging microscopy, Nat. Commun. 15(1), 8019 (2024). https://doi.org/10.1038/s41467-024-52333-2. [Google Scholar]
- DM Jameson, JC Croney. Fluorescence polarization: past, present and future. Comb. Chem. High Throughput Screen. 6(3), 167–176 (2003). https://doi.org/10.2174/138620703106298347. [Google Scholar]
- Sahoo H, Förster resonance energy transfer – A spectroscopic nanoruler: Principle and applications, J. Photochem. Photobiol. C 12(1), 20–30 (2011). https://doi.org/10.1016/j.jphotochemrev.2011.05.001. [Google Scholar]
- Li Y, Cu Y, Luo D. Multiplexed detection of pathogen DNA with DNA-based fluorescence nanobarcodes, Nat. Biotechnol. 23(7), 885–889 (2005). https://doi.org/10.1038/nbt1106. [Google Scholar]
- Liu HW, Chen L, Xu C, Li Z, Zhang H, Zhang X, et al., Recent progresses in small-molecule enzymatic fluorescent probes for cancer imaging, Chem. Soc. Rev. 47(18), 7140–7180 (2018). https://doi.org/10.1039/C7CS00862G. [Google Scholar]
- Mitchell KR, Esene JE, Woolley AT, Advances in multiplex electrical and optical detection of biomarkers using microfluidic devices, Anal. Bioanal. Chem. 414(1), 167–180 (2022). https://doi.org/10.1007/s00216-021-03553-8. [Google Scholar]
- Jiao Y, Du C, Zong L, Guo X, Han Y, Zhang X, et al., 3D vertical-flow paper-based device for simultaneous detection of multiple cancer biomarkers by fluorescent immunoassay, Sens. Actuators B Chem. 306, 127239 (2020). https://doi.org/10.1016/j.snb.2019.127239. [Google Scholar]
- Dysli C, Wolf S, Berezin MY, Sauer L, Hammer M, Zinkernagel MS, Fluorescence lifetime imaging ophthalmoscopy, Prog. Retinal Eye Res. 60, 120–143 (2017). https://doi.org/10.1016/j.preteyeres.2017.06.005. [Google Scholar]
- Witte S, The endoendothelial lining as studied by a fluorescent labeling technique in situ, Thromb. Res. 29(supp-S2), 93–104 (1983). https://doi.org/10.1016/0049-3848(83)90444-9. [Google Scholar]
- Fei X, Gu Y, Progress in modifications and applications of fluorescent dye probe, Prog. Nat. Sci. 19(1), 1–7 (2009). https://doi.org/10.1016/j.pnsc.2008.06.004. [Google Scholar]
- Schneider AFL, Hackenberger CPR, Fluorescent labelling in living cells, Curr. Opin. Biotech. 48, 61–68 (2017). https://doi.org/10.1016/j.copbio.2017.03.012. [Google Scholar]
- Qiao L, Shao X, Gao S, Ming Z, Fu X, Wei Q, Research on endoplasmic reticulum–targeting fluorescent probes and endoplasmic reticulum stress–mediated nanoanticancer strategies: a review, Colloid Surface B 208, 112046 (2021). https://doi.org/10.1016/j.colsurfb.2021.112046. [Google Scholar]
- Mehta VN, Desai ML, Basu H, Singhal RK, Kailasa SK, Recent developments on fluorescent hybrid nanomaterials for metal ions sensing and bioimaging applications: A review, J. Mol. Liq. 333, 115950 (2021). https://doi.org/10.1016/j.molliq.2021.115950. [Google Scholar]
- Zhou H, Tourkakis G, Shi D, Kim DM, Zhang H, Du T, et al., Cell-free measurements of brightness of fluorescently labeled antibodies, Sci. Rep. 7, 41819 (2017). https://doi.org/10.1038/srep41819. [Google Scholar]
- Kong WC, Li CC, Zhang XL, Cui JL, Li XL, Mengy YL, et al., Fluorescent assay for the determination of enrofloxacin using an aptamer and SYBR Green I, Anal. Lett. 58(15), 2654–2666 (2025). https://doi.org/10.1080/00032719.2024.2431984. [Google Scholar]
- Rosa KMD, Washio NMN, Quadros SD, Dreyer JP, Machado VG, Roehrs JA, et al., Mono-and diselenylated fluorescein derivatives: Synthesis using arylseleninic acids as selenylating agents and application as fluorescent probes, J. Mol. Liq. 416, 126528 (2024). https://doi.org/10.1016/j.molliq.2024.126528. [Google Scholar]
- Wang J, Pan X, Zhang K, Zhang S, Sun P, Shi J, et al., Probes based on fluorescein derivatives for rapid detection of H3N2 virus spike proteins without separation, Dyes Pigm. 113080 (2025). https://doi.org/10.1016/j.dyepig.2025.113080. [Google Scholar]
- Carreon JR, Mahon KP, Kelley SO, Thiazole orange-peptide conjugates: Sensitivity of DNA binding to chemical structure, Org. Lett. 6(4), 517–519 (2004). https://doi.org/10.1021/ol0362818. [Google Scholar]
- Wu ZL, Luo A, Grill A, Lao T, Zou Y, Chen Y, Fluorescent detection of O-GlcNAc via tandem glycan labeling, Bioconjugate Chem. 31(9), 2098–2102 (2020). https://doi.org/10.1021/acs.bioconjchem.0c00454. [Google Scholar]
- Mousseau F, Berret J-F, Oikonomou EK, Design and applications of a fluorescent labeling technique for lipid and surfactant preformed vesicles, ACS Omega 4(6), 10485–10493 (2019). https://doi.org/10.1021/acsomega.9b01094. [Google Scholar]
- Cha M, Jeong SH, Bae S, Park JH, Baeg Y, Han DW, et al., Efficient labeling of vesicles with lipophilic fluorescent dyes via the salt-change method, Anal. Chem. 95(14), 5843–5849 (2023). https://doi.org/10.1021/acs.analchem.2c05166. [Google Scholar]
- Xia T, Li N, Fang X, Single-molecule fluorescence imaging in living cells, Annu. Rev. Phys. Chem. 64(1), 459–480 (2013). https://doi.org/10.1146/annurev-physchem-040412-110127. [Google Scholar]
- Liu X, Lin D, Becker W, Niu J, Yu B, Liu L, et al., Fast fluorescence lifetime imaging techniques: A review on challenge and development, J. Innovative Opt. Health Sci. 12(5), 1930003 (2019). https://doi.org/10.1142/S1793545819300039. [Google Scholar]
- Becker W, Fluorescence lifetime imaging–techniques and applications, J. Microsc. 247(2), 119–136 (2012). https://doi.org/10.1111/j.1365-2818.2012.03618.x. [Google Scholar]
- Borst JW, Visser A, Fluorescence lifetime imaging microscopy in life sciences, Meas. Sci. Technol. 21(10), 102002 (2010). https://doi.org/10.1088/0957-0233/21/10/102002. [Google Scholar]
- Torrado B, Pannunzio B, Malacrida L, Digman MA, Fluorescence lifetime imaging microscopy, Nat. Rev. Methods Primers 4(1), 80 (2024). https://doi.org/10.1038/s43586-024-00358-8. [Google Scholar]
- Heikal AA, Intracellular coenzymes as natural biomarkers for metabolic activities and mitochondrial anomalies, Biomarkers Med. 4(2), 241–263 (2010). https://doi.org/10.2217/bmm.10.1. [Google Scholar]
- Ruedas-Rama MJ, Alvarez-Pez JM, Crovetto L, Paredes JM, Orte A, FLIM strategies for intracellular sensing: Fluorescence lifetime imaging as a tool to quantify analytes of interest. Advanced Photon Counting: Applications, Methods, Instrumentation (Springer International Publishing, Cham, 2014), pp. 191–223. https://doi.org/10.1007/4243_2014_67. [Google Scholar]
- Shirshin EA, Shirmanova MV, Gayer AV, Lukina MM, Nikonova EE, Yakimov BP, et al., Label-free sensing of cells with fluorescence lifetime imaging: the quest for metabolic heterogeneity, Proc. Natl. Acad. Sci. 119(9), e2118241119 (2022). https://doi.org/10.1073/pnas.2118241119. [Google Scholar]
- Summers PA, Lewis BW, Gonzalez-Garcia J, Porreca RM, Lim AH, Cadinu P, et al., Visualising G-quadruplex DNA dynamics in live cells by fluorescence lifetime imaging microscopy, Nat. Commun. 12(1), 162 (2021). https://doi.org/10.1038/s41467-020-20414-7. [Google Scholar]
- Pacheco-Liñán PJ, Bravo I, Nueda ML, Albaladejo J, Garzón-Ruiz A. Functionalized CdSe/ZnS Quantum Dots for Intracellular pH Measurements by Fluorescence Lifetime Imaging Microscopy, ACS Sens. 5(7), 2106–2117 (2020). https://doi.org/10.1021/acssensors.0c00719. [Google Scholar]
- Herrera-Ochoa D, Pacheco-Liñán PJ, Bravo I, Garzón-Ruiz A, A novel quantum dot-based pH probe for long-term fluorescence lifetime imaging microscopy experiments in living cells, ACS Appl. Mater. Interface. 14(2), 2578–2586 (2022). https://doi.org/10.1021/acsami.1c19926. [Google Scholar]
- Kanno H, Hiramatsu K, Mikami H, Nakayashiki A, Yamashita S, Nagai A, et al., High-throughput fluorescence lifetime imaging flow cytometry, Nat. Commun. 15, 7376 (2024). https://doi.org/10.1038/s41467-024-51125-y. [Google Scholar]
- Kanno H, Hiramatsu K, Mikami H, Nakayashiki A, Yamashita S, Nagai A, et al., Publisher Correction: High-throughput fluorescence lifetime imaging flow cytometry. Nat. Commun. 16, 582 (2025). https://doi.org/10.1038/s41467-025-55961-4. [Google Scholar]
- Baud D, Gubler DJ, Schaub B, Lanteri MC, Musso D, An update on Zika virus infection, Lancet 390(10107), 2099–2109 (2017). https://doi.org/10.1016/S0140-6736(17)31450-2. [Google Scholar]
- O’Toole RF, The interface between COVID-19 and bacterial healthcare-associated infections, Clin. Microbiol. Infect. 27(12), 1772–1776 (2021). https://doi.org/10.1016/j.cmi.2021.06.001. [Google Scholar]
- Zhang Y, Tang H, Chen W, Zhang J, Nanomaterials used in fluorescence polarization based biosensors, Int. J. Mol. Sci. 23(15), 8625 (2022). https://doi.org/10.3390/ijms23158625. [Google Scholar]
- Camacho R, Täuber D, Scheblykin IG, Fluorescence anisotropy reloaded – emerging polarization microscopy methods for assessing chromophores’ organization and excitation energy transfer in single molecules, particles, films, and beyond, Adv. Mater. 31(22), 1805671 (2019). https://doi.org/10.1002/adma.201805671. [Google Scholar]
- Wan T, Aleem AR, Huang S, Chen R, Zhou Z, Sun M, et al., Autofluorescence free functionalized upconversion nanoparticles-based turn-on aptasensor for highly sensitive and selective sensing of antibiotics, Mater. Today. Adv. 17, 100326 (2023). https://doi.org/10.1016/j.mtadv.2022.100326. [Google Scholar]
- Lee CY, Jung J, Fluorescence polarization assays: Another mainstream in detection of biohazards, Microchem. J. 113523 (2025). https://doi.org/10.1016/j.microc.2025.113523. [Google Scholar]
- Jolley ME, Nasir MS, The use of fluorescence polarization assays for the detection of infectious diseases, Comb. Chem. High. T. Scr. 6(3), 235–244 (2003). https://doi.org/10.2174/138620703106298419. [Google Scholar]
- Rossi AM, Taylor CW, Analysis of protein-ligand interactions by fluorescence polarization, Nat Protoc. 6(3), 365–387 (2011). https://doi.org/10.1038/nprot.2011.305. [Google Scholar]
- Fábio Henrique dos Santos Rodrigues, Delgado GG, Costa TSD, Tasic L, Applications of fluorescence spectroscopy in protein conformational changes and intermolecular contacts, BBA Adv. 3(c), 100091 (2023). https://doi.org/10.1016/j.bbadva.2023.100091. [Google Scholar]
- Sem DS, McNeeley PA, Application of fluorescence polarization to the steady-state enzyme kinetic analysis of calpain II, FEBS Lett. 443(1), 17–19 (1999). https://doi.org/10.1016/s0014-5793(98)01655-x. [Google Scholar]
- Holdgate GA, Hemsley PE, Ligand discovery: High-throughput binding: fluorescence polarization (anisotropy), Methods Mol. Biol. 2263, 231–246 (2021). https://doi.org/10.1007/978-1-0716-1197-5_10. [Google Scholar]
- Li Y, Kang T, Park HG, One-pot, ultrasensitive, and multiplex detection of SARS-CoV-2 genes utilizing self-priming hairpin-mediated isothermal amplification, Biosens. Bioelectron. 237, 115522 (2023). https://doi.org/10.1016/j.bios.2023.115522. [Google Scholar]
- Dutta A, Szekely Z, Guven H, Li XP, McLaughlin JE, Tumer NE, A fluorescence anisotropy-based competition assay to identify inhibitors against ricin and Shiga toxin ribosome interactions, Anal. Biochem. 692, 115580 (2024). https://doi.org/10.1016/j.ab.2024.115580. [Google Scholar]
- Raysyan A, Moerer R, Coesfeld B, Eremin SA, Schneider RJ, Fluorescence polarization immunoassay for the determination of diclofenac in wastewater, Anal. Bioanal. Chem. 413(4), 999–1007 (2021). https://doi.org/10.1007/s00216-020-03058-w. [Google Scholar]
- Duan C, Zhang H, Zhang Y, Li Q, Li P, Mari GM, et al., A Robust homogeneous fluorescence polarization immunoassay for rapid determination of erythromycin in milk, Foods 12(8), 1581 (2023). https://doi.org/10.3390/foods12081581. [Google Scholar]
- Huang J, Chen Y, Sun Z, Gu Y, Li J, One-step platform for Maduramicin and Salinomycin detection based on bispecific monoclonal antibody and interpretation of molecular recognition mechanism, J. Agric. Food Chem. 47, 18557–18565 (2023). https://doi.org/10.1021/acs.jafc.3c06731. [Google Scholar]
- Mukhametova LI, Eremin SA, Fluorescence polarization assays for organic compounds in food safety, Front. Biosci. (Elite Ed) 16(1), 4 (2024). https://doi.org/10.31083/j.fbe1601004. [Google Scholar]
- Förster T, Zwischenmolekulare energiewanderung und fluoreszenz, Ann. Phys. 437, 55–75 (1948). https://doi.org/10.1002/andp.19484370105. [Google Scholar]
- Förster T, Transfer mechanisms of electronic excitation energy, Radiat. Res. Suppl. 2, 326–339 (1960). https://doi.org/10.2307/3583604. [Google Scholar]
- Zhang B, Lyu G, Kelly EA, Evans RC, Förster resonance energy transfer in luminescent solar concentrators, Adv. Sci. 9(23), 2201160 (2022). https://doi.org/10.1002/advs.202201160. [CrossRef] [Google Scholar]
- Stryer L, Haugland RP, Energy transfer: A spectroscopic ruler, Proc. Natl. Acad. Sci. 58(2), 719–726 (1967). https://doi.org/10.1073/pnas.58.2.719. [Google Scholar]
- Nichols JW, Pagano RE, Resonance energy transfer assay of protein-mediated lipid transfer between vesicles, J. Biol. Chem. 258(9), 5368–5371 (1983). https://doi.org/10.1016/S0021-9258(20)81898-5. [Google Scholar]
- Ma L, Yang F, Zheng J, Application of fluorescence resonance energy transfer in protein studies, J. Mol. Struct. 1077, 87–100 (2014). https://doi.org/10.1016/j.molstruc.2013.12.071. [Google Scholar]
- Rudkouskaya A, Sinsuebphon N, Ochoa M, Chen SJ, Mazurkiewicz JE, Intes X, et al., Multiplexed non-invasive tumor imaging of glucose metabolism and receptor-ligand engagement using dark quencher FRET acceptor, Theranostics 10(22), 10309 (2020). https://doi.org/10.7150/thno.45825. [Google Scholar]
- Su B, Zhang Z, Sun Z, Tang Z, Xie X, Chen Q, et al., Hammock BD, Fluonanobody-based nanosensor via fluorescence resonance energy transfer for ultrasensitive detection of ochratoxin A, J. Hazard. Mater. 422, 126838 (2022). https://doi.org/10.1016/j.jhazmat.2021.126838. [Google Scholar]
- Wang Y, Zhang G, Xiao X, Shu X, Fei D, Guang Y, et al., High-performance fluorescent microspheres based on fluorescence resonance energy transfer mode for lateral flow immunoassays, Anal. Chem. 95(48), 17860–17867 (2023). https://doi.org/10.1021/acs.analchem.3c03986. [Google Scholar]
- Qi S, Dong X, Hamed EM, Jiang H, Cao W, Li SFY, Wang Z, Ratiometric fluorescence aptasensor of allergen protein based on multivalent aptamer-encoded dna flowers as fluorescence resonance energy transfer platform, Anal. Chem. 96(18), 11 (2024). https://doi.org/10.1021/acs.analchem.3c05894. [Google Scholar]
All Figures
 |
Fig. 1 Energy levels and radiative transition from the excited state (S1/S2) to the ground state (S0). |
| In the text | |
 |
Fig. 2 (a) Fluorescent detection of O‑GlcNAc via Tandem Glycan Labeling. (b) Fluorescent labeling technique for lipid and surfactant preformed vesicles. (c) Efficient labeling of vesicles with lipophilic fluorescent dyes via the Salt-Change method. |
| In the text | |
 |
Fig. 3 (a) Visualising G-quadruplex DNA dynamics in live cells by fluorescence lifetime imaging microscopy. (b) Functionalized CdSe/ZnS quantum dots for intracellular pH measurements by fluorescence lifetime imaging microscopy. (c) A novel quantum dot-based pH probe for long-term fluorescence lifetime imaging microscopy experiments in living cells. (d) High-throughput fluorescence lifetime imaging flow cytometry. |
| In the text | |
 |
Fig. 4 Schematic diagram of fluorescence polarization principle. |
| In the text | |
 |
Fig. 5 (a) Schematic diagram of fluorescence polarization immunoassay for determining diclofenac in wastewater. (b) Schematic diagram of homogeneous fluorescence polarization immunoassay for rapid determination of erythromycin in milk. (c) Schematic diagram of one-step FPIA detection of maduramycin and salinomycin based on bispecific monoclonal antibodies. |
| In the text | |
 |
Fig. 6 Schematic diagram of fluorescence resonance energy transfer. |
| In the text | |
 |
Fig. 7 (a) Schematic illustration of non-invasive tumor imaging based on multi parameter glucose metabolism and receptor-ligand engagement using dark quenched FRET acceptor. (b) Schematic illustration of Fluonanobody-based nanosensor via fluorescence resonance energy transfer for ultrasensitive detection of ochratoxin A. (c) Schematic diagram of high-performance fluorescent microspheres based on fluorescence resonance energy transfer mode for lateral flow immunoassay. (d) Schematic illustration of the Principle of the Ratiometric Fluorescence Aptasensor Based on Multivalent Aptamer Encoded DNA Flowers. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


